Michael T. Sandholzer PhD
I translate complex multi-omics and clinical trial data into actionable insights for immuno-oncology research, from single-cell sequencing to biomarker strategy.
5+
Years Experience
7+
Projects Led
6
Publications

I translate complex multi-omics and clinical trial data into actionable insights for immuno-oncology research, from single-cell sequencing to biomarker strategy.
5+
Years Experience
7+
Projects Led
6
Publications

I am a computational biologist specializing in cancer immunology and multi-omics data science, with hands-on experience across the full data lifecycle: from wet-lab experimental design to data creation, multi-omics integration, and biologically grounded interpretation.
In my current postdoctoral role at the University Hospital of Basel, I act as scientific lead for 7+ multi-omics projects, integrating single-cell transcriptomics, immune repertoire profiling, and perturbation data from longitudinal clinical and pre-clinical studies. Beyond executing analyses, I scope projects with clinicians and experimental teams, translate biological questions into data science strategies, and communicate results in decision-oriented formats that directly inform biomarker prioritization and R&D direction.
At Novartis, working in GMP-regulated cell therapy manufacturing taught me that scientific rigor must align with business needs and regulatory constraints — a mindset that stays with me in every collaborative setting. My formal training at ETH Zurich (MSc Chemistry) and LMU Munich (BSc Chemistry & Biochemistry) gave me a mechanistic perspective I bring to every omics dataset.
I am particularly drawn to roles where scientific depth, stakeholder interaction, and real-world clinical impact intersect: in biopharma R&D, scientific consulting, or translational research.
Capabilities that emerge from working at the intersection of biology, computation, and clinical research, and that are rarely found combined in a single person.
Integrating computational analysis with biological interpretation: every model output is evaluated in the context of what is biologically plausible.
Bridging clinicians, wet-lab scientists, and bioinformaticians, translating between domains so that every stakeholder understands what the data mean for their decisions.
Translating high-dimensional multi-omics datasets into decision-ready results, moving from terabytes of sequencing data to a concrete, actionable biological conclusion.
Understanding how experimental data are generated and where artifacts arise, enabling more accurate interpretation and preventing misleading computational conclusions.
Recognizing meaningful biological signals within complex datasets, distinguishing true biology from technical noise, batch effects, and experimental confounders.
Selected research projects combining multi-omics, clinical trial data, and computational methods to advance cancer immunotherapy.
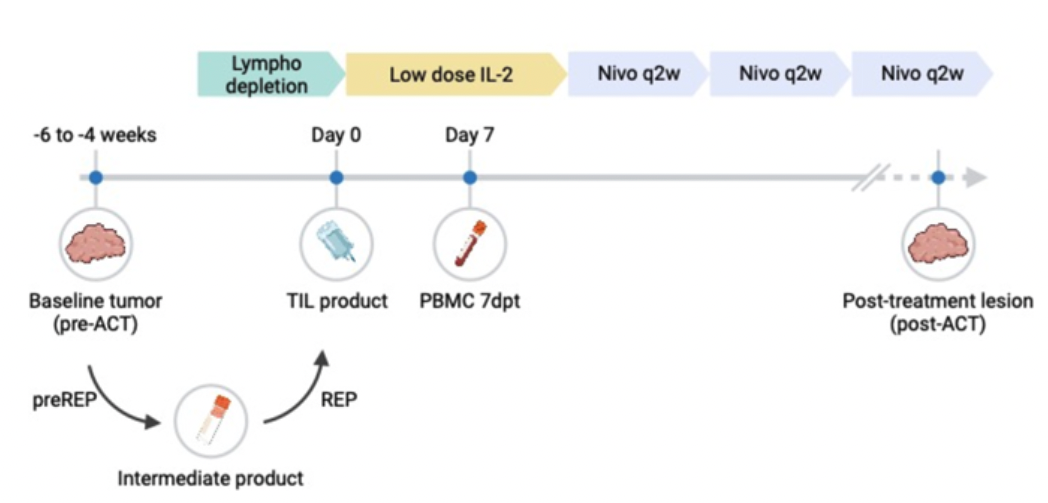
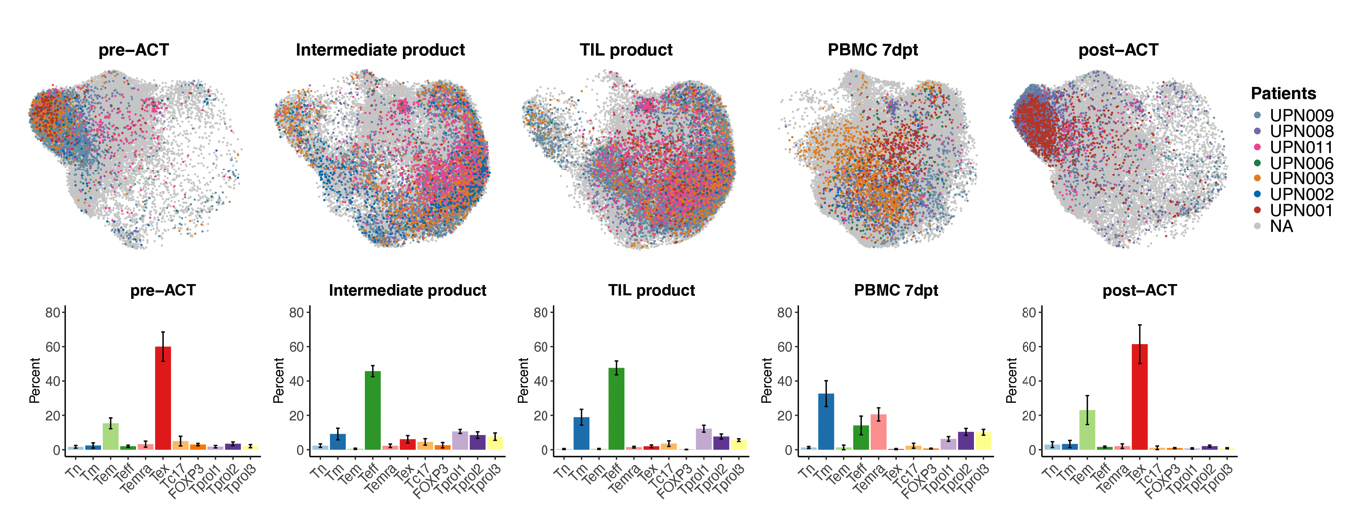
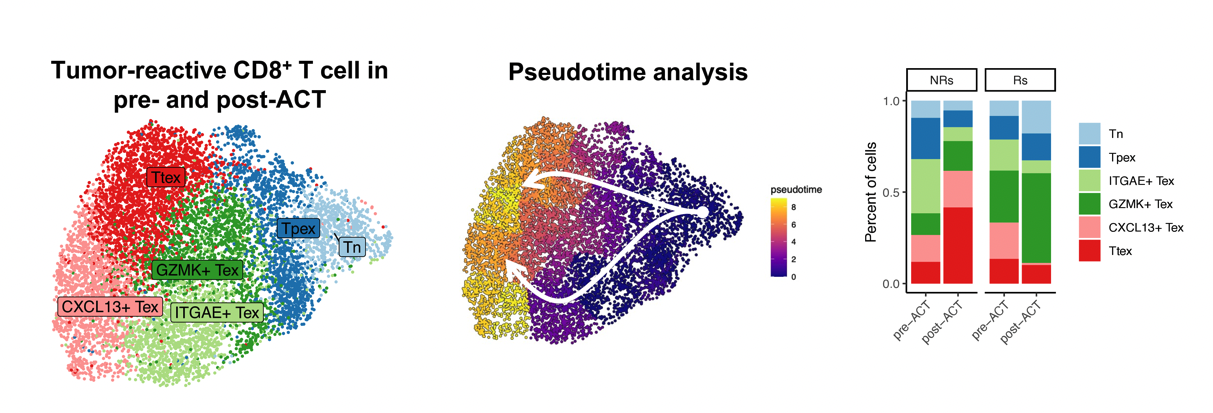
Used single-cell RNA and TCR sequencing to profile T cell dynamics across seven patients in a Phase I/II TIL therapy clinical trial. We found that tumor-reactive T cells are reinvigorated during ex vivo expansion, yet reacquire exhaustion signatures after infusion into the tumor microenvironment — providing a mechanistic basis for response and relapse in adoptive cell therapy.
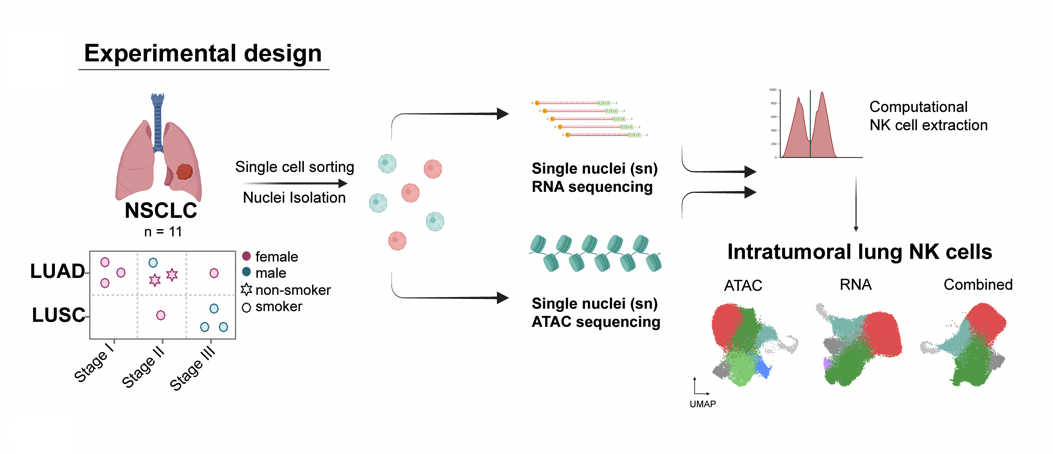
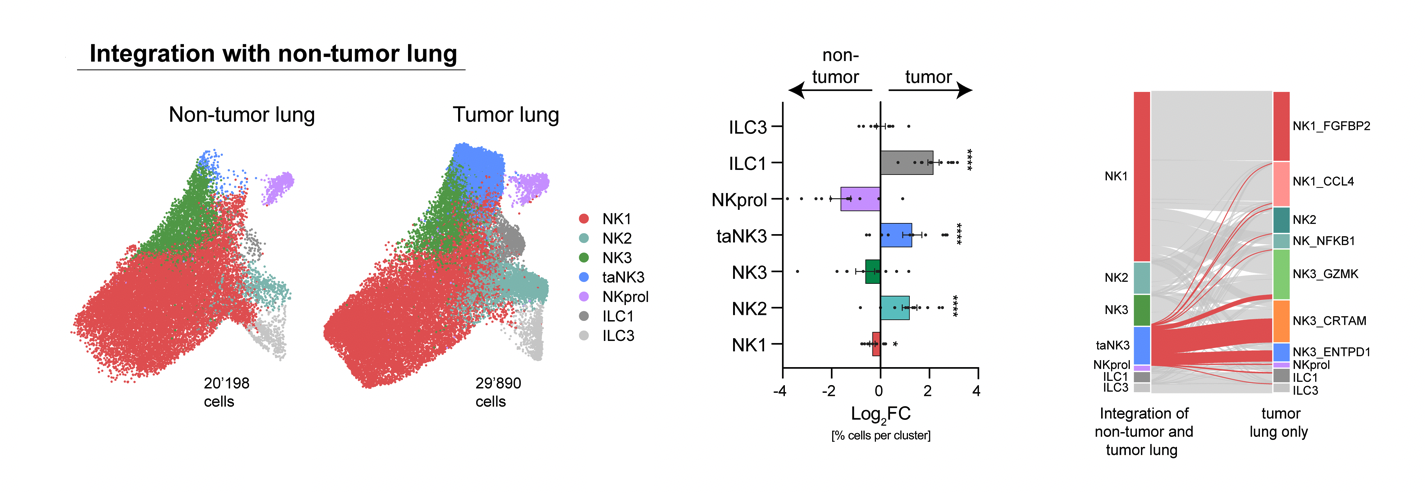
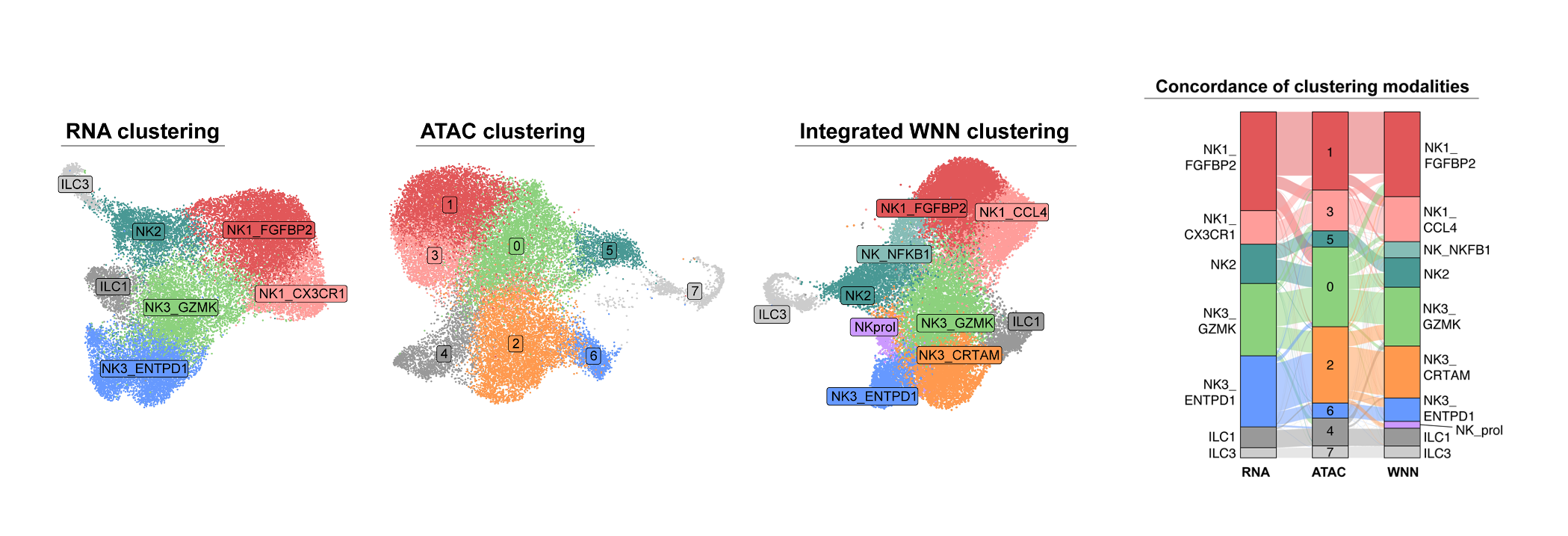
Resolved NK cell heterogeneity in non-small cell lung cancer using paired snRNA-seq and snATAC-seq from 11 treatment-naïve patients. Identified CD39+ tumor-associated NK cells as a distinct effector subset characterized by activation of interferon-stimulated genes and a cytotoxic gene program, providing mechanistic insight and a novel cellular target for NK cell-based immunotherapy.
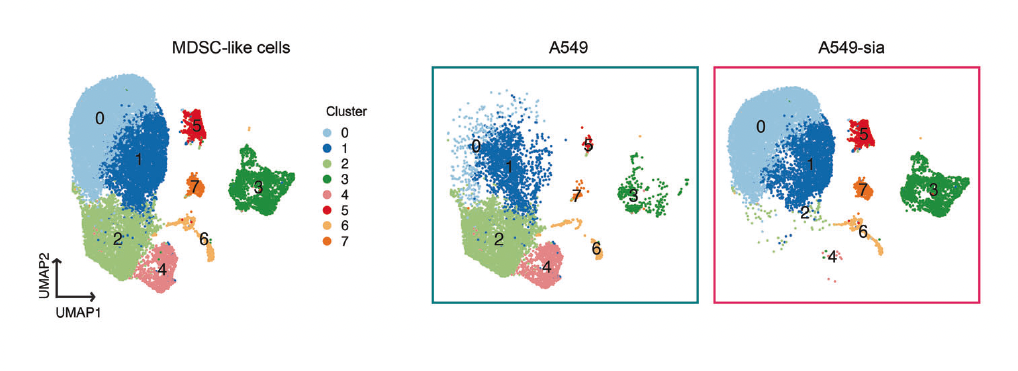
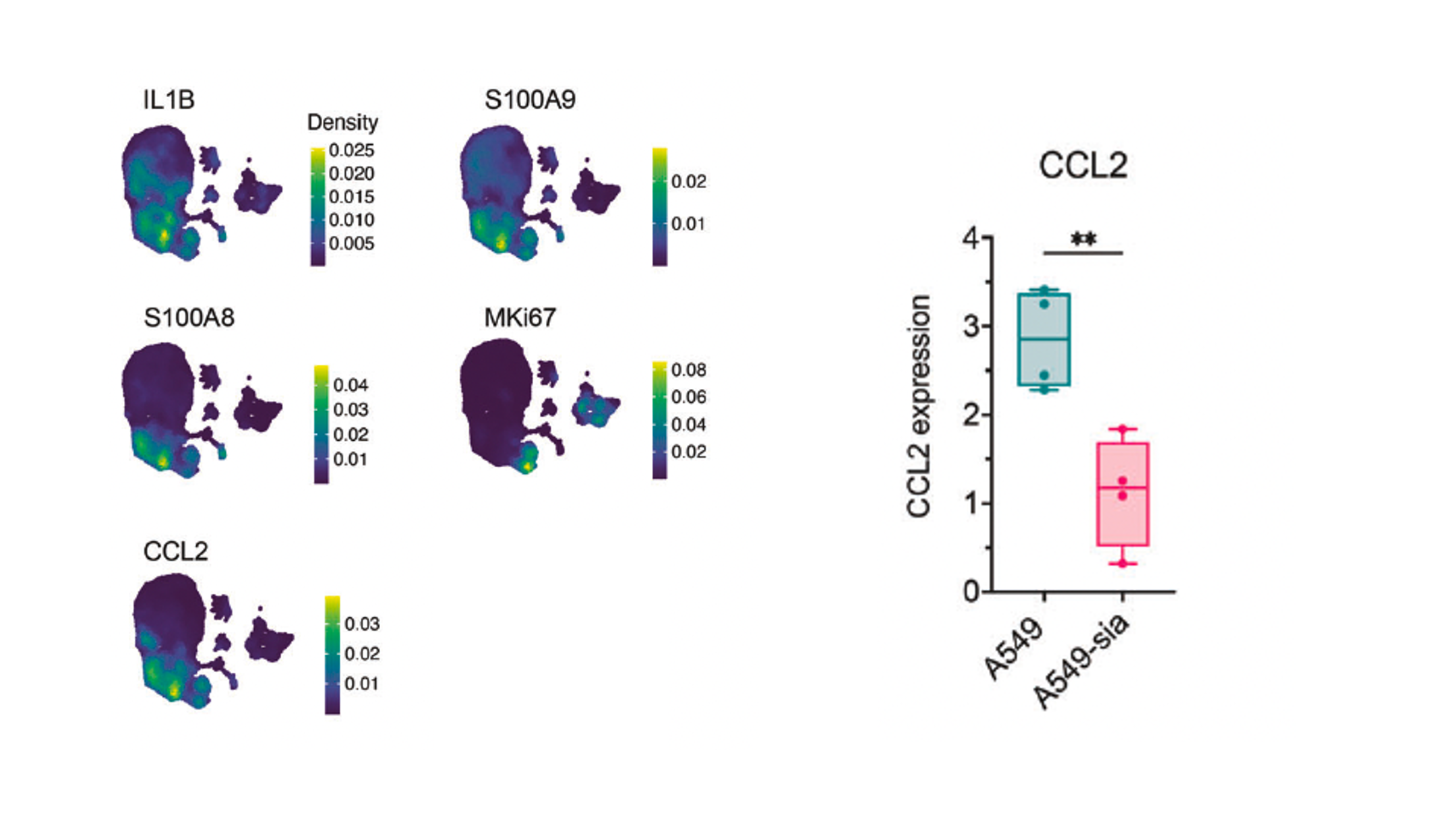
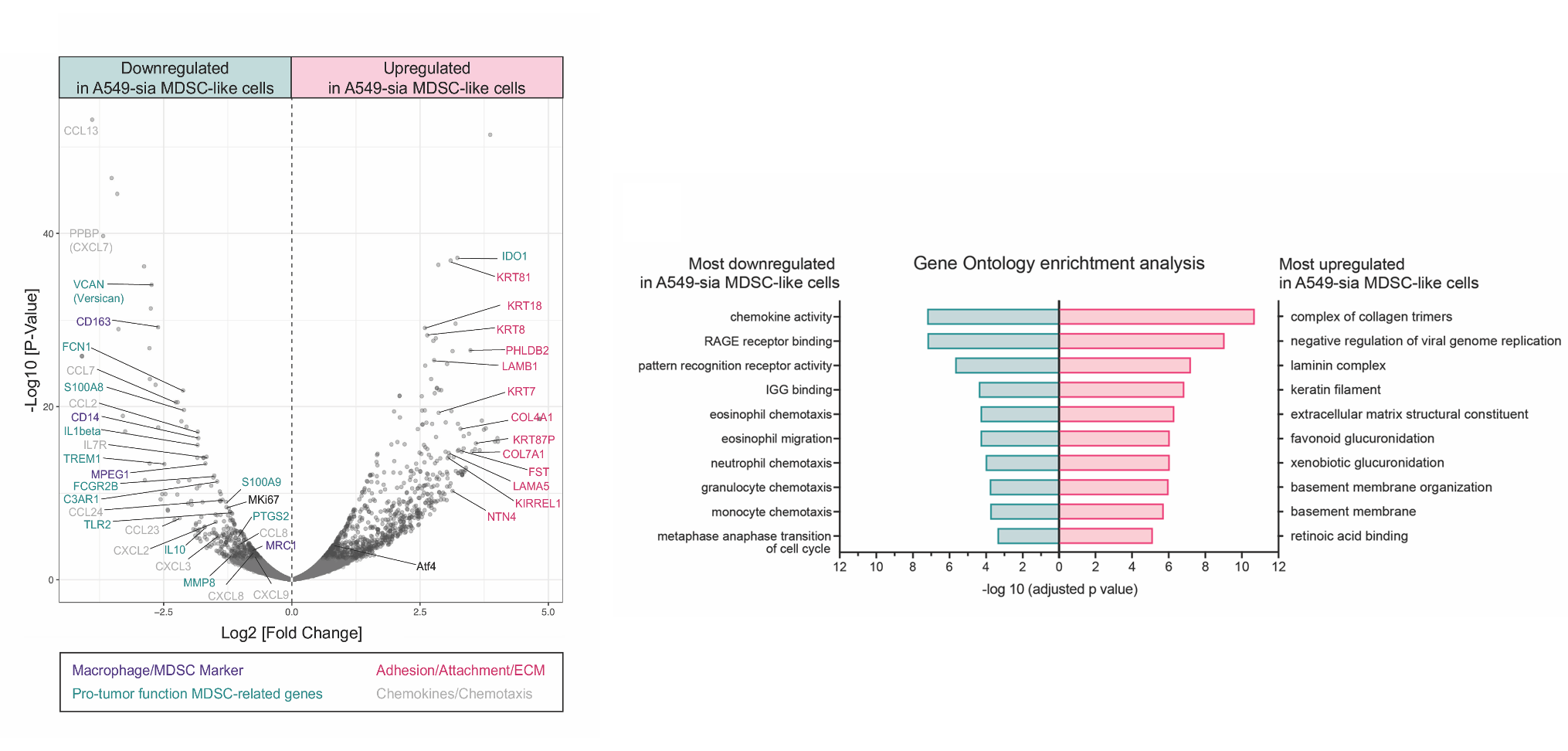
Investigated how sialylated glycans on cancer cell surfaces drive immune suppression in the tumor microenvironment. Siglec receptor engagement on myeloid-derived suppressor cells induced CCL2 secretion, suppressing T cell infiltration. CRISPR knockout of Siglec-E in myeloid cells improved tumor control in vivo, identifying the glycan–Siglec axis as a tractable immunotherapy target.
University Hospital of Basel · Basel, CH
University Hospital of Basel · Basel, CH
Novartis Pharma Stein AG · Stein, CH
University of Basel · Basel, CH
Swiss Federal Institute of Technology (ETH Zurich) · Zurich, CH
Ludwig Maximilian University (LMU) · Munich, DE
* These authors contributed equally to this work.
Sandholzer, M. T., et al. Longitudinal profiling of tumor-reactive T cells during TIL therapy in metastatic melanoma. bioRxiv, 2025.09.23.678066 (2025).
Serger, C., Rebuffet, L.*, Sandholzer, M. T., et al. Integrated single cell analysis identifies CD39+ tumor-associated NK cells with cytotoxic potential in lung cancer. bioRxiv, 2025.08.26.672316 (2025).
König, D.*, Sandholzer, M. T.*, et al. Melanoma clonal heterogeneity leads to secondary resistance after adoptive cell therapy with tumor-infiltrating lymphocytes. Cancer Immunol Res, 12, 814–821 (2024).
Wieboldt, R., Sandholzer, M., et al. Engagement of sialylated glycans with Siglec receptors on suppressive myeloid cells inhibits anti-cancer immunity via CCL2. Cell Mol Immunol, 21, 495–509 (2024).
König, D., Kasenda, B., Sandholzer, M., et al. Adoptive cell therapy with tumor-infiltrating lymphocytes in combination with nivolumab in patients with advanced melanoma. Immunooncol Technol, 24, 100728 (2024).
Pellegrino, C., Favalli, N., Sandholzer, M., et al. Impact of Ligand Size and Conjugation Chemistry on the Performance of Universal Chimeric Antigen Receptor T-Cells for Tumor Killing. Bioconjug Chem (2020).
Fusi, I., Serger, C., Herzig, P., Germann, M., Sandholzer, M. T., et al. PD-1–targeted cis-delivery of an IL-2 variant induces a multifaceted antitumoral T cell response in human lung cancer. Sci Transl Med, 17, eadr3718 (2025).
Professional references from academic, clinical, and industry supervisors are available upon request. Please reach out via email or LinkedIn.
mt.sandholzer@gmail.com
linkedin.com/in/sandholzer
GitHub
github.com/Sandholzer
Google Scholar
scholar.google.com
ORCID
0009-0005-4842-6013
Curriculum Vitae
Download PDF
Open to opportunities
I am interested in roles at the intersection of data science, precision medicine, and biomedical research: in biopharma R&D, scientific consulting, bioinformatics platforms, or translational research teams. Feel free to reach out.